Advance fundamental understanding of genome biology and develop the genome-scale engineering technologies needed to design, build, and control plants and microbes for desired beneficial purposes.

Metabolic Mixology. Cell-free prototyping offers a metabolic mixology approach for accelerating the design of biosynthetic pathways for sustainable manufacturing. [Reprinted by permission from Springer Nature from Karim, A. S., et al. 2020. “In Vitro Prototyping and Rapid Optimization of Biosynthetic Enzymes for Cell Design,” Nature Chemical Biology 16(8), 912–19. DOI: 10.1038/s41589-020-0559-0. Copyright 2020.]
The ability to deliberately manipulate and design biological systems has large implications for Genomic Science Program (GSP) research, including the elucidation of gene function at the organismal level; production of novel, high-value fuels, chemicals, and materials; and manipulation of plant and microbial behavior and interactions for the development of sustainable biofuel and bioproduct production systems. The program seeks to enable a future in which biological systems can be designed for specific purposes in silico and be built and tested using automated procedures, delivering new biosystems for deployment into the bioeconomy. Given the complexity and breadth of the requisite scientific challenges, this goal will require a broad, interdisciplinary, and cross-institutional approach.
GSP therefore recognizes that novel approaches to organizing scientific interactions and collaborations, such as virtual laboratories, may offer significant advantages. The program will also work to leverage the many synergies of current biosystems design developments with other Biological and Environmental Research (BER) program elements into a diverse research portfolio in biological engineering. As the promise of synthetic biology begins to bear fruit, the biotechnology industry is expected to play a growing role in the U.S. economy. BER is poised to contribute to growing opportunities in this area through its support of basic biosystems design research relevant to DOE’s mission space.
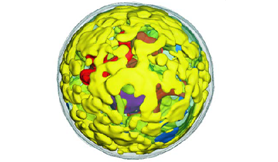
Gene Targets. Cryo-soft x-ray tomography shows substantial accumulation of starch (yellow) in chloroplasts of C. zofingiensis as part of research on the alga’s physiological and genetic response to changes in glucose availability. This will help identify target genes for developing new engineered strains that accumulate high amounts of biofuels and bioproducts. [From Roth, M. S., et al. 2019. “Regulation of Oxygenic Photosynthesis during Trophic Transitions in the Green Alga Chromochloris zofingiensis.” The Plant Cell, 31(3), 579–601. DOI: 10.1105/tpc.18.00742. Cryo-soft X-ray tomography capabilities supported by DOE Office of Basic Energy Sciences.]
- Discover and develop novel platform organisms across a range of physiologies as chassis for synthetic biology.
- Develop innovative genome-engineering tools, including large-scale DNA synthesis and intracellular delivery, recoded and minimal genomes, orthogonal pathways, and cell-free systems.
- Develop high-throughput genome editing, automated screening, characterization, phenotyping, and testing of engineered organisms.
- Elucidate gene function at the genome scale to develop generalized approaches for biological engineering.
- Provide computer-aided design tools, including artificial intelligence and machine-learning techniques, in an integrated, open-access platform for plants and microbes as part of an in silico or virtual laboratory capability.
- Develop analytical and characterization tools to understand processes of organic and inorganic synthesis and degradation in engineered organisms.
- Elucidate mechanisms for the acquisition, storage, transport, and chemical transformation of substrates in engineered organisms.
- Support research to build new genetic, regulatory, and biosynthetic networks to biologically produce useful molecules and materials that may or may not exist in nature.
- Develop new approaches for novel macromolecular design, characterization, and testing.
- Expand the “design-build-test-learn” cycle to organismal consortia.
