09/18/2020
New Approach Helps Determine How Much Microbial Community Composition Is Driven by Selection and How Much by Chance
A novel computational framework allows researchers to quantify the relative importance of different ecological processes in the composition of microbiomes
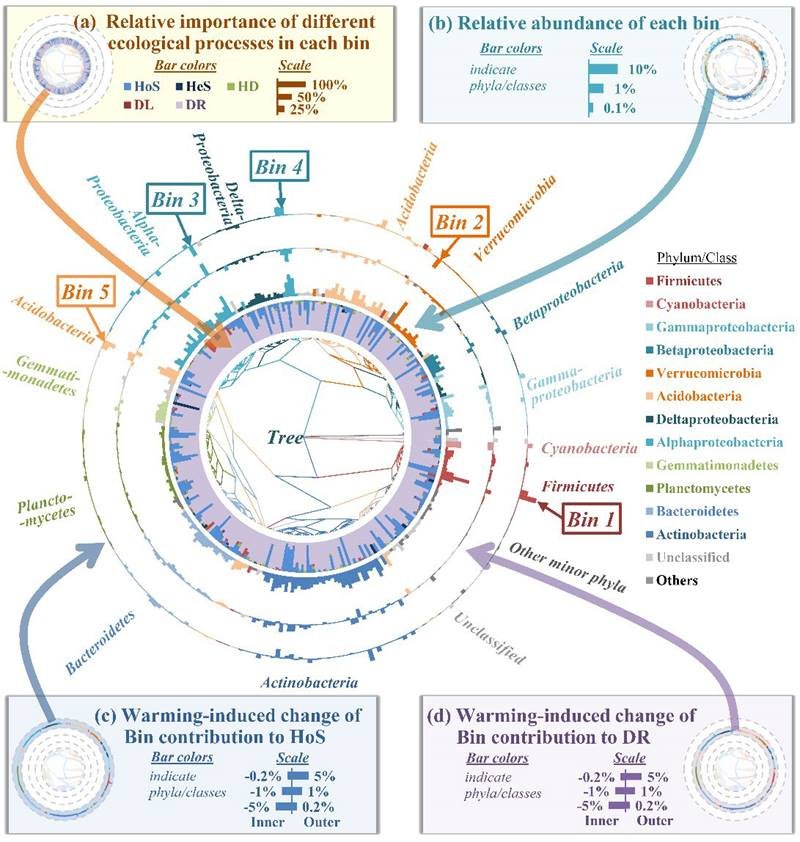
The new iCAMP framework can provide insights into assembly mechanisms of different phylogenetic groups, such as the relative importance of selection (HoS/HeS), dispersal (HD/DL), and drift in shaping variation of each ‘bin’ under different conditions.
[Ning, D., et al., A quantitative framework reveals ecological drivers of grassland microbial community assembly in response to warming. Nature Communications 11, 4717 (2020). [DOI: 10.1038/s41467-020-18560-z. CC BY 4.0.]
The Science
Life on our planet consists of many communities of plants, animals, and microbes. These communities are shaped by complex ecological processes. Those processes include natural selection, migration, and random shifts in births and deaths. Quantifying the relative importance of these processes is a major challenge in ecology research. The challenge is especially great for the ecology of microbes. This study develops an approach named iCAMP. This approach is based on the concept that different processes can govern different groups of species in a diverse community. iCAMP showed excellent performance on simulated microbial communities. When applied to real-world grassland microbial communities, iCAMP revealed that environmental changes altered the relative importance of the ecological processes.
The Impact
Sorting out the factors that control biodiversity is a crucial but difficult task. This is particularly true for below-ground microbial communities. Cutting-edge genomic technologies allow scientists to easily detect millions of genes and genomes. This creates a new challenge: how to use data to reveal the ecological mechanisms that affect the diversity of microbes in a community. This study developed an effective and robust new tool called iCAMP. The tool quantitatively assesses the ecological processes that shape diversity and change in microbial communities, or microbiomes. The tool works at the level of individual species or groups of organisms rather than at the level of entire communities.
Summary
Selection, dispersal, diversification, and drift are major processes affecting the assembly of microbial communities, but defining their relative importance is challenging. In this study, researchers developed a framework to quantitatively infer community assembly mechanisms. This framework, called phylogenetic bin-based null model analysis (iCAMP), integrates a series of computational and statistical approaches to analyze the effects of ecological processes on individual lineages within a given microbial community. On simulated communities, iCAMP showed substantially higher accuracy, precision, sensitivity, and specificity than entire community-based approaches.
Applying iCAMP to subsurface microbial communities in a grassland under experimental warming revealed that homogeneous selection and drift were the dominant ecological processes shaping this microbial community. Using data collected during multiple years, iCAMP captured a decrease of drift and an increase of homogeneous selection over time, primarily imposed on the group Bacillales. In addition, iCAMP enabled the researchers to reveal drought and plant productivity as major drivers behind the variation of homogeneous selection.
Principal Investigator
Jizhong Zhou
University of Oklahoma
jzhou@ou.edu
BER Program Manager
Boris Wawrik
U.S. Department of Energy, Biological and Environmental Research (SC-33)
Biological Systems Science Division
boris.wawrik@science.doe.gov
Funding
This work was supported by the Department of Energy Office of Science, Office of Biological and Environmental Research, the Ecosystems and Networks Integrated with Genes and Molecular Assemblies (ENIGMA) Scientific Focus Area at Lawrence Berkeley National Laboratory, and by the National Science Foundation.
References
Ning, D., et al., A quantitative framework reveals ecological drivers of grassland microbial community assembly in response to warming. Nature Communications 11, 4717 (2020). [DOI: 10.1038/s41467-020-18560-z
