Conformational Equilibria Underlying Electron Bifurcation and Transfer in Thermotoga Maritima Fix/EtfABCX: Novel Structural States and Their Correlation to Catalysis by Anaerobic SEC-SAXS
Authors:
Daniel T. Murray1* (murraydt@lbl.gov), Xiaoxuan Ge2, Gerrit J. Schut2, Michal Hammel1, Michael W. W. Adams2, and Greg L. Hura1
Institutions:
1Lawrence Berkeley National Laboratory; and 2University of Georgia
Goals
The conformational states associated with electron bifurcation in metalloenzyme complexes (bifurcases) is sought in order to obtain a mechanistic understanding of their functions. Static structures of bifurcase enzymes often lack the detail necessary to fully provide such an understanding, so a small-angle X-ray scattering (SAXS) approach has been adopted for the characterization pipeline. Anaerobic size-exclusion chromatography-coupled SAXS (SEC-SAXS) achieves sufficient isolation of macromolecular species and protection from oxygen to provide an assay for bifurcase and metalloenzyme complex structure in solution. It is with these tools that the current project seeks to understand electron bifurcation for the potential application of engineering biofuel systems.
Abstract
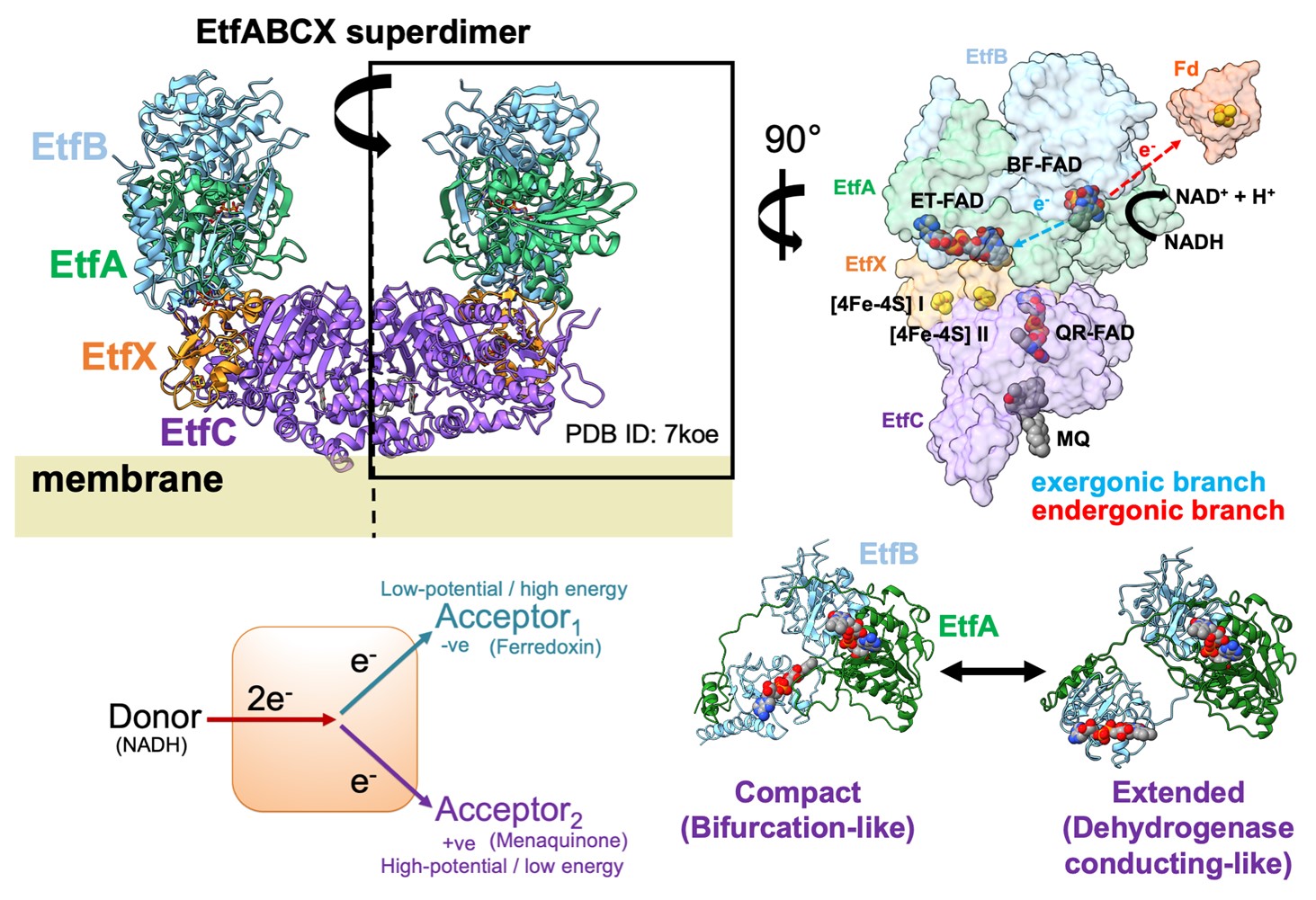
Electron bifurcation is a newly discovered albeit evolutionarily ancient means of energy conservation used by anaerobic microorganisms that simultaneously delivers low- and high-potential electrons to acceptors in a thermodynamically favorable fashion. The cryo-EM structure of one enzyme performing flavin-based bifurcation from Thermotoga maritima–Fix/EtfABCX–was recently solved to 2.9 Å resolution (Feng et al. 2021). This structure suggested a model for its catalytic mechanism that implies the low-potential electrons generated during bifurcation transfer to ferredoxin and high-potential electrons reduce menaquinone. This structure depicts Fix/EtfABCX as a symmetric superdimer, each half of the superdimer formed by an ABCX heterotetramer, and as membrane-associated. Presented here, anaerobic small-angle X-ray scattering analysis (Classen et al. 2013; Rosenberg, Hura, and Hammel 2022) on Fix/EtfABCX shows the enzyme to adopt an asymmetric morphology in solution, which can be further exaggerated or reverted towards symmetry by the presence of NAD+ or NADH, respectively. These conditions also impact the overall rigidity of the system which, together with the observed conformational changes, imply coenzyme presence primes the system for accepting reducing equivalents, interaction with substrates, and progression through its catalytic cycle. Interestingly, the presence of NADH induces formation of supertetrameric Fix/EtfABCX from two superdimer particles, suggesting change in redox state and/or presence of this coenzyme imparts a conformation conducive to oligomerization and a subsequent role in the bifurcation process. Collectively, these solution state observations provide structural narratives for which the states of Fix/EtfABCX must abide during its catalysis and also serve as a foundation for future studies of oxygen-sensitive metalloenzymes involved in electron bifurcation.
Image
References
Feng, X., et al. 2021. “Cryoelectron Microscopy Structure and Mechanism of the Membrane-Associated Electron-Bifurcating Flavoprotein Fix/EtfABCX.” Proceedings of the National Academy of Sciences 118, e2016978118. DOI:10.1073/pnas.2016978118.
Classen, S., et al. 2013. “Implementation and Performance of SIBYLS: a Dual Endstation Small-Angle X-Ray Scattering and Macromolecular Crystallography Beamline at the Advanced Light Source.” Journal of Applied Crystallography 46, 1–13. DOI: 10.1107/s0021889812048698.
Rosenberg, D. J., G. L. Hura, and M. Hammel 2022. “Size Exclusion Chromatography Coupled Small Angle X-Ray Scattering with Tandem Multiangle Light Scattering at the SIBYLS Beamline”. Methods Enzymology 677, 191–219. DOI: 10.1016/bs.mie.2022.08.031.
Funding Information
This work was conducted at the Advanced Light Source (ALS), a national user facility operated by Lawrence Berkeley National Laboratory on behalf of the Department of Energy, Office of Basic Energy Sciences, through the Integrated Diffraction Analysis Technologies (IDAT) program, supported by DOE Office of Biological and Environmental Research.
